Two brain maps look correlated. But the brain is spatially autocorrelated—nearby regions tend to have similar values simply because they are neighbours. A standard correlation test ignores this structure, which inflates p-values and produces false positives.
neuromapr exists to answer one question honestly: is the similarity between two brain maps more than spatial structure alone would predict?
It generates surrogate maps that preserve the spatial autocorrelation of the original data, then measures where the real correlation falls in that null distribution. The framework follows Markello et al. (2022).
Comparing two brain maps
The core workflow: load data, compare, interpret. We simulate two brain maps here to keep things self-contained, but in practice you would read real data from GIFTI or NIfTI files.
set.seed(42)
n <- 100
coords <- matrix(rnorm(n * 3), ncol = 3)
distmat <- as.matrix(dist(coords))
map_a <- rnorm(n)
map_b <- 0.4 * map_a + rnorm(n, sd = 0.8)Two maps with a planted correlation around 0.4, plus a distance matrix describing the spatial layout. The simplest comparison:
result <- compare_maps(map_a, map_b, verbose = FALSE)
result
#>
#> ── Brain Map Comparison
#> Method: pearson
#> r = 0.5302, p = 1.4e-08
#> n = 100The r and parametric p tell the familiar
story. But that p assumes independence between
observations—an assumption the brain violates everywhere.
Adding a spatial null model
To get an honest p-value, we generate surrogate versions of
map_a that preserve its spatial autocorrelation, then check
whether the observed correlation is unusual against correlations with
those surrogates.
neuromapr provides eight null model methods. For parcellated data
with a distance matrix, variogram matching (burt2020) is a
strong default:
result_null <- compare_maps(
map_a, map_b,
null_method = "burt2020",
distmat = distmat,
n_perm = 500L,
seed = 1,
verbose = FALSE
)
#> Generating burt2020 nulls ■■■■■■■■■■■■■■■■■ 52% | ETA: 4s
#> Generating burt2020 nulls ■■■■■■■■■■■■■■■■■■■■■■■■■■ 84% | ETA: 2s
result_null
#>
#> ── Brain Map Comparison
#> Method: pearson
#> r = 0.5302, p = 1.4e-08
#> n = 100
#> ────────────────────────────────────────────────────────────────────────────────
#> Null model: burt2020 (500 permutations)
#> p_null = 0.002The p_null value now accounts for spatial
autocorrelation. If it remains small, the relationship between the two
maps is unlikely to be an artifact of shared spatial structure.
Visualising the null distribution
The comparison object stores all null correlations, making it straightforward to see where the observed value falls:
null_df <- data.frame(r = result_null$null_r)
obs_r <- result_null$r
ggplot2::ggplot(null_df, ggplot2::aes(x = r)) +
ggplot2::geom_histogram(
bins = 30, fill = "steelblue", alpha = 0.7
) +
ggplot2::geom_vline(
xintercept = obs_r,
linetype = "dashed", color = "red", linewidth = 1
) +
ggplot2::labs(
x = "Null correlation (r)",
y = "Count"
) +
ggplot2::theme_minimal()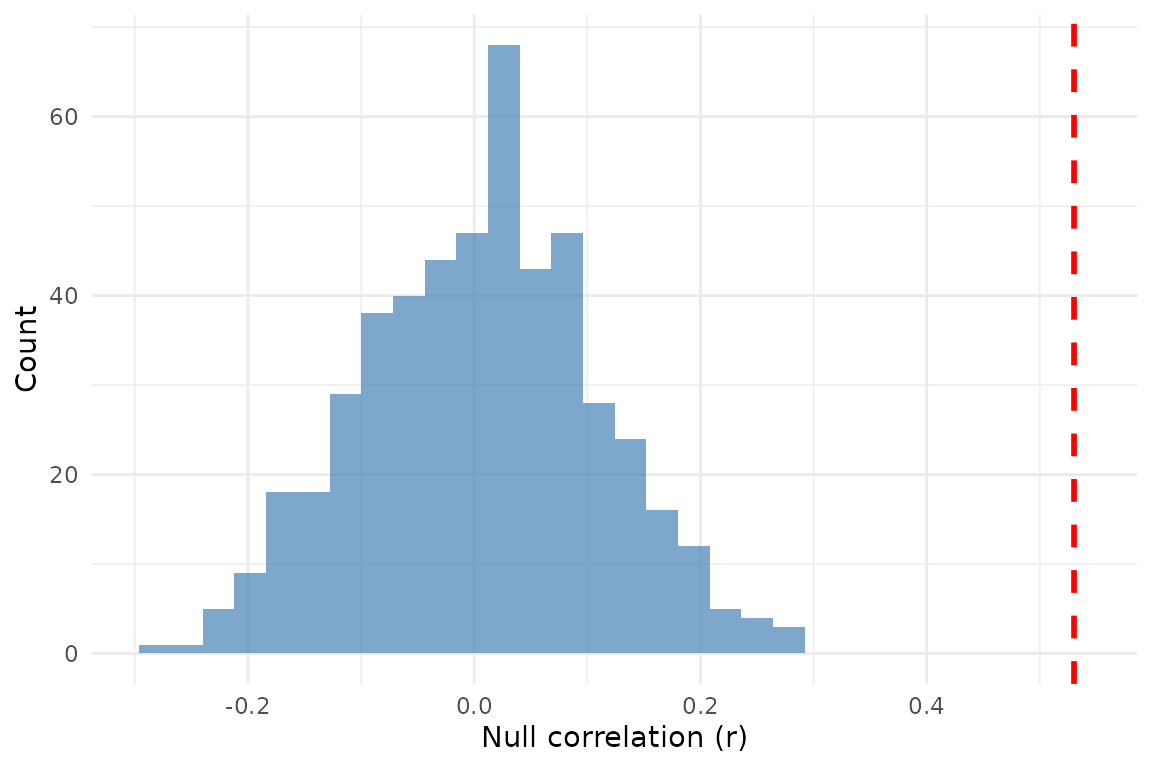
Null correlations from 500 variogram-matching surrogates. The dashed red line marks the observed correlation.
When the observed correlation sits well outside the null distribution, the relationship is more likely genuine. When it sits inside, spatial autocorrelation alone can explain the similarity.
Pre-computing null distributions
Generating nulls is the expensive part. If you want to compare the same map against several targets, generate once and reuse:
nulls <- generate_nulls(
map_a,
method = "moran",
distmat = distmat,
n_perm = 200L,
seed = 42
)
nulls
#>
#> ── Null Distribution
#> • Method: moran
#> • Permutations: 200
#> • Observations: 100The null_distribution object has print,
summary, plot, and as.matrix
methods. The summary gives per-element null means and
standard deviations. The plot method shows the null
distribution for a chosen element:
plot(nulls, parcel = 1L)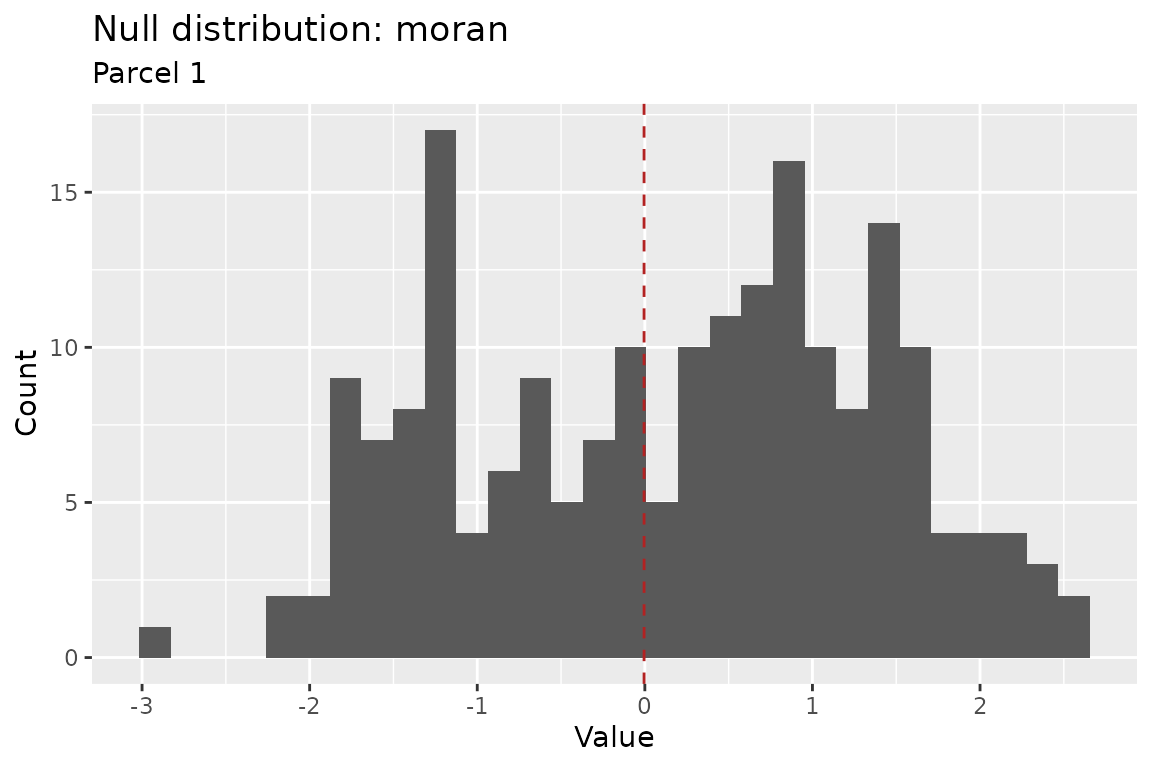
Built-in plot method for a null_distribution object.
Pass the pre-computed object to compare_maps():
compare_maps(map_a, map_b, nulls = nulls, verbose = FALSE)
#>
#> ── Brain Map Comparison
#> Method: pearson
#> r = 0.5302, p = 1.4e-08
#> n = 100
#> ────────────────────────────────────────────────────────────────────────────────
#> Null model: moran (200 permutations)
#> p_null = 0.00498No redundant computation when the same source map appears in multiple comparisons.
Working with real brain map files
In practice, brain maps live on disk. compare_maps() and
read_brain_map_values() accept file paths directly:
values <- read_brain_map_values("cortical_thickness.func.gii")
result <- compare_maps(
"path/to/map_a.func.gii",
"path/to/map_b.func.gii",
null_method = "moran",
distmat = distmat
)Both GIFTI (.func.gii) and NIfTI (.nii.gz)
formats are supported. No separate reading step needed.
Accessing the neuromaps atlas collection
The neuromaps project curates a registry of published brain map annotations—PET receptor maps, gene expression, cortical thickness, and more. neuromapr provides direct access to this collection:
neuromaps_available()
neuromaps_available(source = "beliveau", tags = "pet")Once you find the annotation you want, download it:
paths <- fetch_neuromaps_annotation(
"abagen", "genepc1", "fsaverage",
density = "10k"
)Files are cached locally, so subsequent calls skip the download. The registry is fetched from the neuromaps GitHub repository and cached for the session.
Custom metrics with permtest_metric()
compare_maps() is built around correlation. For other
metrics—mean absolute error, cosine similarity, anything that takes two
vectors and returns a scalar—permtest_metric() provides the
same null model machinery:
mae <- function(a, b) mean(abs(a - b))
result_mae <- permtest_metric(
map_a, rnorm(n),
metric_func = mae,
n_perm = 200L,
seed = 1
)
result_mae$observed
#> [1] 1.117007
result_mae$p_value
#> [1] 0.6517413Add null_method for spatially-constrained surrogates
instead of random permutation:
result_spatial <- permtest_metric(
map_a, rnorm(n),
metric_func = mae,
n_perm = 200L,
seed = 1,
null_method = "burt2020",
distmat = distmat
)
#> Generating burt2020 nulls ■■■■■■■■■■■■■■■■■■■■■■■ 74% | ETA: 1s
result_spatial$p_value
#> [1] 0.8159204Choosing a null model
The right model depends on your data:
| Situation | Recommended method |
|---|---|
| Parcellated data with distance matrix |
"burt2020" or "moran"
|
| Vertex-level with sphere coordinates |
"spin_hungarian" or "alexander_bloch"
|
| Parcellated with sphere coordinates |
"baum" or "cornblath"
|
| Spatial autoregressive structure | "burt2018" |
vignette("null-models") walks through each method in
detail.
What comes next
This vignette covered the essential workflow: compare maps, generate spatially-constrained null distributions, and interpret results. From here:
-
vignette("null-models")dives into all eight null model methods and their assumptions. -
vignette("parcellation")covers aggregating vertex data into parcels and parcel-level null models. -
vignette("surface-geometry")explains geodesic distances, surface graphs, and vertex area calculations.
References
Markello RD, Hansen JY, Liu Z-Q, et al. (2022). neuromaps: structural and functional interpretation of brain maps. Nature Methods, 19, 1472–1480. doi:10.1038/s41592-022-01625-w